node2vec_online (Enterprise)
From Memgraph 3.0, dynamic graph algorithms became a part of the Memgraph Enterprise license. To use dynamic graph algorithms, you need to enable Memgraph Enterprise license. If you want to know more and learn how this affects you, read our announcement.
The node2vec_online algorithm learns and updates temporal node embeddings on
the fly for tracking and measuring node similarity over time in graph streams.
The algorithm creates similar embeddings for two nodes (e.g. v and u) if there
is an option to reach one node from the other across edges that appeared
recently. In other words, the embedding of a node v should be more similar to
the embedding of node u if we can reach u by taking steps backward to node
v across edges that appeared before the previous one. These steps backward
from one node to the other form a temporal walk. It is temporal since it depends
on when the edge appeared in the graph.
To make two nodes more similar and to create these temporal walks, the Node2Vec Online algorithm uses the StreamWalk updater and Word2Vec learner.
StreamWalk updater is a machine for sampling temporal walks. A sampling of the
walk is done in a backward fashion because we look only at the incoming edges of
the node. Since one node can have multiple incoming edges, when sampling a walk,
StreamWalk updater uses probabilities to determine which incoming edge of the
node it will take next, and that way leading to a new node. These probabilities
are computed after the edge arrives and before temporal walk sampling.
Probability represents a sum over all temporal walks z ending in node v
using edges arriving no later than the latest one of already sampled ones in the
temporal walk. When the algorithm decides which edge to take next for temporal
walk creation, it uses these computed weights (probabilities). Every time a new
edge appears in the graph, these probabilities are updated just for two nodes of
a new edge.
After walks sampling, Word2Vec learner uses these prepared temporal walks to
make node embeddings more similar using the gensim Word2Vec module. These
sampled walks are given as sentences to the gensim Word2Vec module, which then
optimizes for the similarity of the node embeddings in the walk with stochastic
gradient descent using a skip-gram model or continuous-bag-of-words (CBOW).
Embeddings capture the graph topology, relationships between nodes, and further relevant information. How the embeddings capture this inherent information of the graph is not fixed.
Capturing information in networks often shuttles between two kinds of similarities: homophily and structural equivalence. Under the homophily hypothesis, nodes that are highly interconnected and belong to similar network clusters or communities should be embedded closely together. In contrast, under the structural equivalence hypothesis, nodes that have similar structural roles in networks should be embedded closely together (e.g., nodes that act as hubs of their corresponding communities).
Currently, our implementation captures for homophily - nodes that are highly interconnected and belong to similar network clusters or communities.
Check out the Node embeddings in dynamic graphs by Ferenc Béres, Róbert Pálovics, Domokos Miklós Kelen and András A. Benczúr.
| Trait | Value |
|---|---|
| Module type | module |
| Implementation | Python |
| Graph direction | directed |
| Edge weights | unweighted |
| Parallelism | sequential |
Too slow?
If this algorithm implementation is too slow for your use case, open an issue on Memgraph’s GitHub repository and request a rewrite to C++!
Procedures
The basic node classification workflow is as follows:
- Load data to Memgraph.
- Set updater parameters by calling the
set_streamwalk_updater()procedure. - Set learner parameters by calling the
set_streamwalk_learner()procedure. - Call the
update()procedure to set the node embeddings. - Inspect the results by calling the
get()procedures. - If you want to reset the parameters and start again, call the
restart()procedure.
You can execute this algorithm on graph projections, subgraphs or portions of the graph.
set_streamwalk_updater()
Use the procedure to sample the temporal random walks in the graph. Once the
set_word2vec_learner() parameters are set, you first need to call the
reset() procedure to update the parameters.
Input:
-
subgraph: Graph(OPTIONAL) ➡ A specific subgraph, which is an object of type Graph returned by theproject()function, on which the algorithm is run. If subgraph is not specified, the algorithm is computed on the entire graph by default. -
half_life: integer➡ Half-life [seconds], used in the temporal walk probability calculation. -
max_length: integer➡ The maximum length of the sampled temporal random walks. -
beta: float➡ The damping factor for long paths. -
cutoff: integer➡ A temporal cutoff in seconds to exclude very distant past. -
sampled_walks: integer➡ The number of sampled walks for each edge update. -
full_walks: boolean➡ An option to return every node of the sampled walk for representation learning (full_walks=True) or only the endpoints of the walk (full_walks=False).
Output:
message: string➡ Whether parameters are set or they need to be reset.
Usage:
To start sampling the temporal random walks, use the following query:
CALL node2vec_online.set_streamwalk_updater(7200, 3, 0.9, 604800, 4, False)
YIELD message
RETURN message;set_word2vec_learner()
Use the procedure to calculate embeddings. To recalculate embeddings with
different parameters, you first need to call the reset() procedure to update
the parameters.
Input:
-
subgraph: Graph(OPTIONAL) ➡ A specific subgraph, which is an object of type Graph returned by theproject()function, on which the algorithm is run. If subgraph is not specified, the algorithm is computed on the entire graph by default. -
embedding_dimension: integer➡ The number of dimensions in the representation of the embedding vector. -
learning_rate: float➡ The learning rate. -
skip_gram: boolean (default=True)➡ Whether to use skip-gram model (True) or continuous-bag-of-words (CBOW) (False). -
negative_rate: integer➡ Negative rate for Gensim Word2Vec model. -
threads: integer➡ Maximum number of threads for parallelization.
Output:
message: string➡ Whether parameters are set or they need to be reset.
Usage:
To set the learner parameter values, use the following query:
CALL node2vec_online.set_word2vec_learner(128, 0.01, True, 10, 1)
YIELD message
RETURN message;update()
Use the update() procedure to (re)calculate embeddings.
Input:
-
subgraph: Graph(OPTIONAL) ➡ A specific subgraph, which is an object of type Graph returned by theproject()function, on which the algorithm is run. If subgraph is not specified, the algorithm is computed on the entire graph by default. -
edges: mgp.List[mgp.Edge]➡ A list of relationships added to the graph. Embeddings will be (re)calculated only for the starting and ending nodes of those new relationships.
Usage:
You can use the update() procedure in two ways.
You can create a trigger, so every time a relationships is added to graph, the trigger calls a procedure and makes an update:
CREATE TRIGGER trigger ON --> CREATE BEFORE COMMIT
EXECUTE CALL node2vec_online.update(createdEdges)
RETURN *;Or, you can filter out certain relationships and call the algorithm with those relationships:
MATCH (n)-[e]->(m)
WITH COLLECT(e) as edges
CALL node2vec_online.update(edges)
RETURN edges;get()
The get() procedure returns the node embeddings.
Input:
subgraph: Graph(OPTIONAL) ➡ A specific subgraph, which is an object of type Graph returned by theproject()function, on which the algorithm is run. If subgraph is not specified, the algorithm is computed on the entire graph by default.
Output:
node: mgp.Vertex➡ The node in the graph for which embedding exists.embedding: mgp.List[mgp.Number]➡ Embedding for the given node.
Usage:
To return the calculated node embeddings, use the following query:
CALL node2vec_online.get()
YIELD node, embedding
RETURN node, embedding;reset()
Use the procedure to reset the updater and learner parameters.
Output:
message: string➡ Message that parameters are ready to be set again.
Usage:
To reset the parameters, use the following query:
CALL node2vec_online.reset()
YIELD message
RETURN message;help()
The procedure will return procedures available in the query module, their parameters and descriptions.
Output:
name: string➡ The name of available functions.value: string➡ Documentation for every function.
Usage:
To get help, use the following query:
CALL node2vec_online.help()
YIELD name, value RETURN name, value;Example
Database state
The database contains the following data:
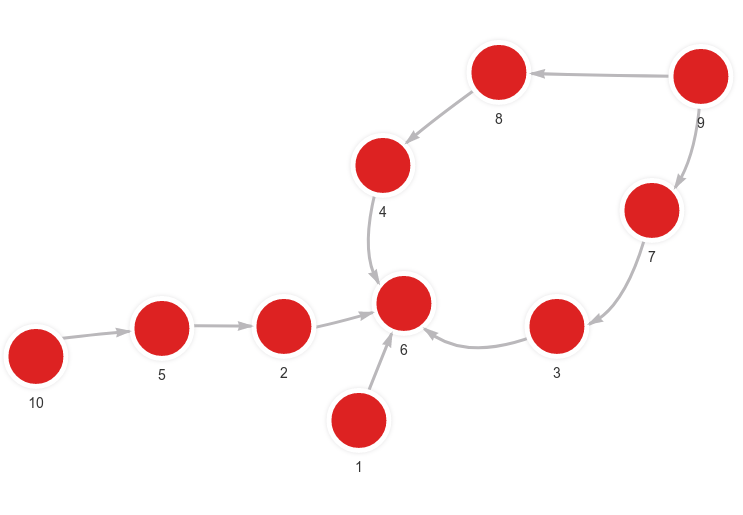
Created with the following Cypher queries:
MERGE (n:Node {id: 1}) MERGE (m:Node {id: 6}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 2}) MERGE (m:Node {id: 6}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 10}) MERGE (m:Node {id: 5}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 5}) MERGE (m:Node {id: 2}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 9}) MERGE (m:Node {id: 7}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 7}) MERGE (m:Node {id: 3}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 3}) MERGE (m:Node {id: 6}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 9}) MERGE (m:Node {id: 8}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 8}) MERGE (m:Node {id: 4}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 4}) MERGE (m:Node {id: 6}) CREATE (n)-[:RELATION]->(m);Set the parameters
To set the parameters for the updater and the learner, use the following query:
CALL node2vec_online.set_streamwalk_updater(7200, 2, 0.9, 604800, 2, True)
YIELD message RETURN message;
CALL node2vec_online.set_word2vec_learner(2, 0.01, True, 1, 1)
YIELD message RETURN message;Set a trigger
In order to recalculate node embeddings when new relationships are added to the graph, create a trigger:
CREATE TRIGGER trigger ON --> CREATE BEFORE COMMIT
EXECUTE CALL node2vec_online.update(createdEdges)
RETURN *;Add new relationships
Use the following queries to add new relationships between nodes:
MERGE (n:Node {id: 1}) MERGE (m:Node {id: 6}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 2}) MERGE (m:Node {id: 6}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 10}) MERGE (m:Node {id: 5}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 5}) MERGE (m:Node {id: 2}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 9}) MERGE (m:Node {id: 7}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 7}) MERGE (m:Node {id: 3}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 3}) MERGE (m:Node {id: 6}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 9}) MERGE (m:Node {id: 8}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 8}) MERGE (m:Node {id: 4}) CREATE (n)-[:RELATION]->(m);
MERGE (n:Node {id: 4}) MERGE (m:Node {id: 6}) CREATE (n)-[:RELATION]->(m);Calculate embeddings
Use the following query to get the calculated embeddings:
CALL node2vec_online.get()
YIELD node, embedding
RETURN node, embedding
ORDER BY node.id;Results:
+-------------------------+-------------------------+
| node | embedding |
+-------------------------+-------------------------+
| (:Node {id: 1}) | [0.255167, 0.450464] |
| (:Node {id: 2}) | [-0.465147, -0.35584] |
| (:Node {id: 3}) | [-0.243008, -0.0908009] |
| (:Node {id: 4}) | [-0.414261, -0.472441] |
| (:Node {id: 5}) | [-0.250771, -0.188169] |
| (:Node {id: 6}) | [-0.0268114, 0.0118215] |
| (:Node {id: 7}) | [-0.226831, 0.327703] |
| (:Node {id: 8}) | [0.143829, 0.0495937] |
| (:Node {id: 9}) | [0.369025, -0.0766736] |
| (:Node {id: 10}) | [0.322944, 0.448649] |
+-------------------------+-------------------------+