
Understanding Community Detection Algorithms With Python NetworkX
Introduction
While humans are very good at detecting distinct or repetitive patterns among a few components, the nature of large interconnected networks makes it practically impossible to perform such basic tasks manually. Groups of densely connected nodes are easy to spot visually, but more sophisticated methods are needed to perform these tasks programmatically. Community detection algorithms are used to find such groups of densely connected components in various networks.
M. Girvan and M. E. J. Newman have proposed one of the most widely adopted community detection algorithms, the Girvan-Newman algorithm. According to them, groups of nodes in a network are tightly connected within communities and loosely connected between communities.
In this article, you will learn the basic principles behind community detection algorithms, their specific implementations, and how you can run them using Python and NetworkX.
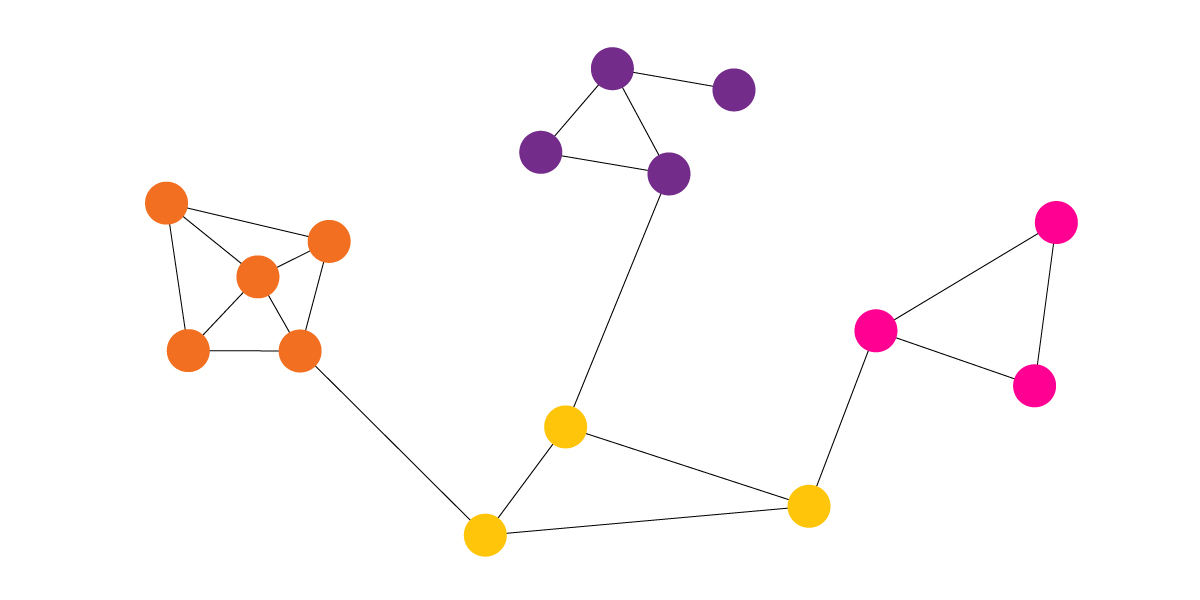
Community Detection Example Applications
Because networks are an integral part of many real-world problems, community detection algorithms have found their way into various fields, ranging from social network analysis to public health initiatives.
- A known use case is the detection of terrorist groups in social networks by tracking their activities and interactions. Similarly, groups of malicious bots can be detected on online social platforms.
- Community detection can be used to study the dynamics of certain groups that are susceptible to epidemic diseases. Other types of diseases can be studied similarly to discover common links among patients.
- One of the most recent use cases, community evolution prediction, involves the prediction of changes in network structures.
Community Detection Techniques
There are two main types of community detection techniques, agglomerative and divisive.
Agglomerative methods generally start with a network that contains only nodes of the original graph. The edges are added one-by-one to the graph, while stronger edges are prioritized over weaker ones. The strength of an edge, or weight, is calculated differently depending on the specific algorithm implementation.
On the other hand, divisive methods rely on the process of removing edges from the original graph iteratively. Stronger edges are removed before weaker ones. At every step, the edge-weight calculation is repeated, since the weight of the remaining edges changes after an edge is removed. After a certain number of steps, we get clusters of densely connected nodes, a.k.a. communities.
Community Detection in NetworkX
- Girvan-Newman algorithm: The Girvan-Newman algorithm detects communities by progressively removing edges from the original network.
- Fluid Communities algorithm: This algorithm is based on the simple idea of fluids interacting in an environment, expanding and pushing each other.
- Label Propagation algorithm: Label propagation is a semi-supervised machine learning algorithm that assigns labels to previously unlabeled data points.
- Clique Percolation algorithm: The algorithm finds k-clique communities in a graph using the percolation method.
- Kernighan-Lin algorithm: This algorithm partitions a network into two sets by iteratively swapping pairs of nodes to reduce the edge cut between the two sets.
Girvan-Newman Algorithm
For the detection and analysis of community structures, the Girvan-Newman algorithm relies on the iterative elimination of edges that have the highest number of shortest paths between nodes passing through them. By removing edges from the graph one-by-one, the network breaks down into smaller pieces, so-called communities. The algorithm was introduced by Michelle Girvan and Mark Newman.
How does it work?
The idea is to find which edges in a network occur most frequently between other pairs of nodes by finding edge betweenness centrailities. The edges joining communities are then expected to have a high edge betweenness. The underlying community structure of the network will be much more fine-grained once the edges with the highest betweenness are eliminated which means that communities will be much easier to spot.
The Girvan-Newman algorithm can be divided into four main steps:
- For every edge in a graph, calculate the edge betweenness centrality.
- Remove the edge with the highest betweenness centrality.
- Calculate the betweenness centrality for every remaining edge.
- Repeat steps 2–4 until there are no more edges left.
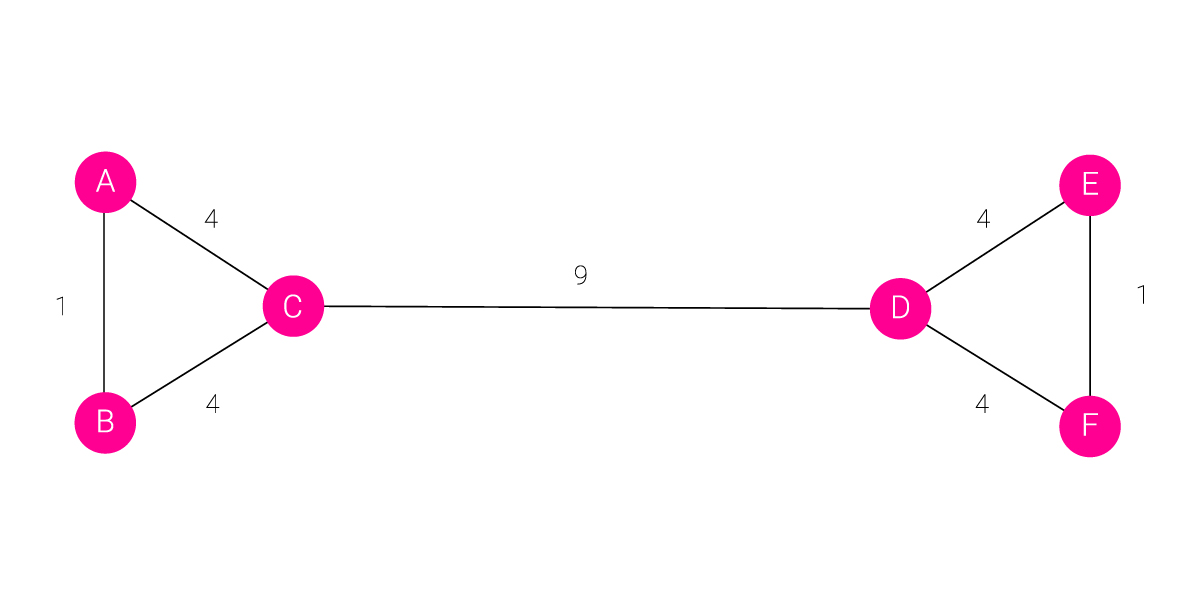
In this example, you can see how a typical graph looks like when edges are assigned weights based on the number of shortest paths passing through them. To keep things simple, we only calculated the number of undirected shortest paths that pass through an edge. The edge between nodes A and B has a strength/weight of 1 because we don’t count A->B and B->A as two different paths.

The Girvan-Newman algorithm would remove the edge between nodes C and D because it is the one with the highest strength. As you can see intuitively, this means that the edge is located between communities. After removing an edge, the betweenness centrality has to be recalculated for every remaining edge. In this example, we have come to the point where every edge has the same betweenness centrality.
Betweenness centrality
Betweenness centrality measures the extent to which a node or edge lies on paths between nodes. Nodes and edges with high betweenness may have considerable influence within a network under their control over information passing between others.
The calculation of betweenness centrality is not standardized and there are many ways to solve it. It is defined as the number of shortest paths in the graph that passes through the node or edge divided by the total number of shortest paths.
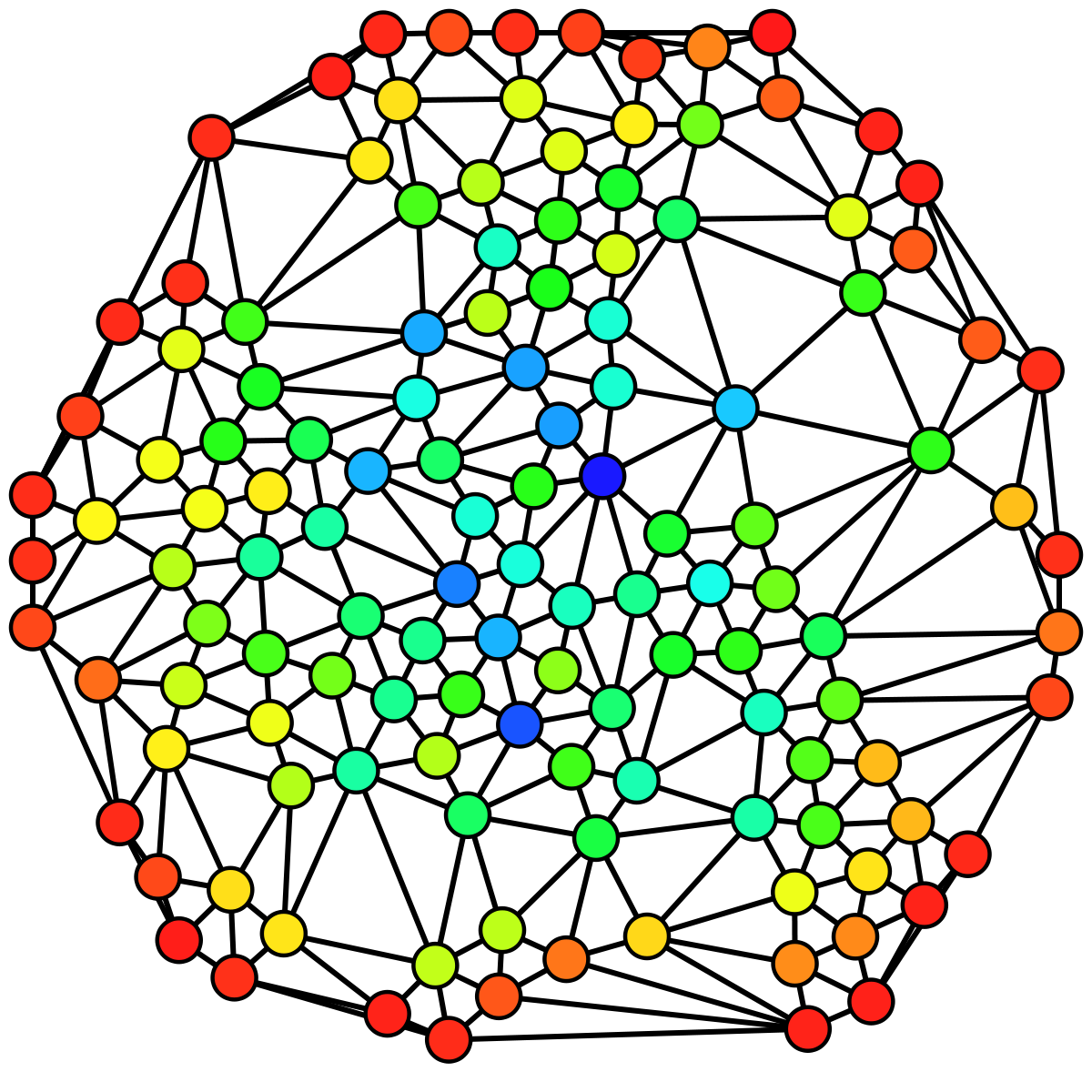
In this example, you can see an undirected graph colored based on the betweenness centrality of each node from lowest (red) to highest (blue).
Pseudocode
In each iteration, calculate the betweenness centrality for all edges in the graph. Remove the edge with the highest centrality. Repeat until there are no more edges left.
REPEAT
LET n BE number of edges in the graph
FOR i=0 to n-1
LET B[i] BE betweenness centrality of edge i
IF B[i] > max_B THEN
max_B = B[i]
max_B_edge = i
ENDIF
ENDFOR
REMOVE edge i FROM graph
UNTIL number of edges in graph is 0
Community Detection Algorithms in NetworkX
girvan_newman(G, most_valuable_edge=None)Method input
The first input parameter of the method, G, is a NetworkX graph.
The second parameter, most_valuable_edge, is a function that takes a graph as input and returns the edge that should be removed from the graph in each iteration. If no function is specified, the edge with the highest betweenness centrality will be chosen in each iteration.
Method output
The output of the method is an iterator over tuples of sets of nodes in G. Each set of nodes represents a community and each tuple is a sequence of communities at a particular level (iteration) of the algorithm.
Example
import matplotlib.pyplot as plt
import networkx as nx
from networkx.algorithms.community.centrality import girvan_newman
G = nx.karate_club_graph()
communities = girvan_newman(G)
node_groups = []
for com in next(communities):
node_groups.append(list(com))
print(node_groups)
color_map = []
for node in G:
if node in node_groups[0]:
color_map.append('blue')
else:
color_map.append('green')
nx.draw(G, node_color=color_map, with_labels=True)
plt.show()The output is:
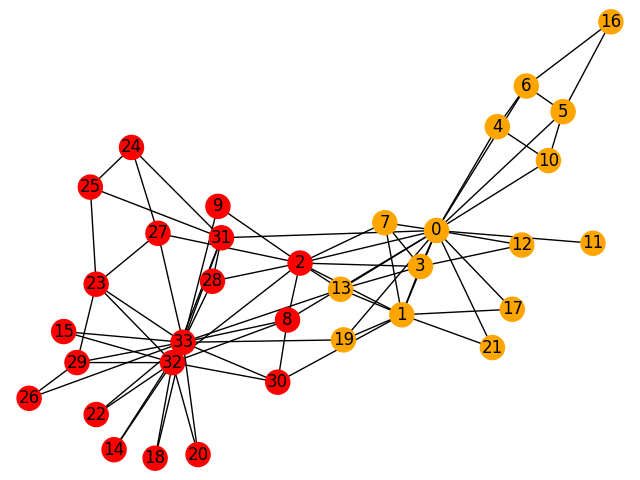
[[0, 1, 3, 4, 5, 6, 7, 10, 11, 12, 13, 16, 17, 19, 21],
[2, 8, 9, 14, 15, 18, 20, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33]]The network has been divided into two distinct communities:
Community Detection in Memgraph
While the NetworkX package may be enough on its own to learn about graph theory and algorithms, in production, we often require a permanent storage solution. Using Memgraph, an in-memory graph database, as the storage solution provides additional benefits and functionalities to NetworkX. In this section, we will focus on how to implement a custom Cypher procedure that uses query modules in Memgraph to perform community detection.
Creating a Custom Query Module
Memgraph supports extending the query language with user-written procedures. These procedures are grouped into modules (Query Modules), which can then be loaded on startup or later on. We are going to create such a procedure to work with the NetworkX package.
To get started, we need to create and mount a volume to access the query_modules directory. This directory contains all of the built-in query modules and it's where you save new custom query modules. Create an empty directory ~modules on your host machine and execute the following command:
docker volume create --driver local --opt type=none --opt device=$(pwd)/modules --opt o=bind modules
Now, you can start Memgraph and mount the created volume:
docker run -it --rm -v modules:/usr/lib/memgraph/query_modules -p 7687:7687 memgraph
Everything from the directory /usr/lib/memgraph/query_modules will be visible/editable in your mounted modules volume and vice versa. If you take a look at the /query_modules directory, you will find built-in query modules that come prepackaged with Memgraph. To learn more about them, visit the How to Use Query Modules Provided by Memgraph? guide.
Create a file called community.py in the modules directory and copy the code below into it.
import mgp
import networkx as nx
from networkx.algorithms import community
from mgp_networkx import MemgraphMultiDiGraph
@mgp.read_proc
def detect(
ctx: mgp.ProcCtx
) -> mgp.Record(communities=mgp.List[mgp.List[mgp.Vertex]]):
networkxGraph = nx.DiGraph(MemgraphMultiDiGraph(ctx=ctx))
communities_generator = community.girvan_newman(networkxGraph)
return mgp.Record(communities=[
list(s) for s in next(communities_generator)])If you are using a Linux machine and don't have the right permissions to create a new file in your modules directory, run the following command:
sudo chown -R <user-name> modules
We just created a new query module with the procedure detect() that utilizes the Girvan–Newman method for finding communities in a graph. Before we can call it, the newly created query module has to be loaded. Execute the following command in Memgraph Lab:
CALL mg.load_all()We need actual data in our database instance to try out the detect() procedure. In Memgraph Lab, navigate to the Datasets tab in the left sidebar and load the Capital cities and borders dataset.
Now, you can call the procedure with:
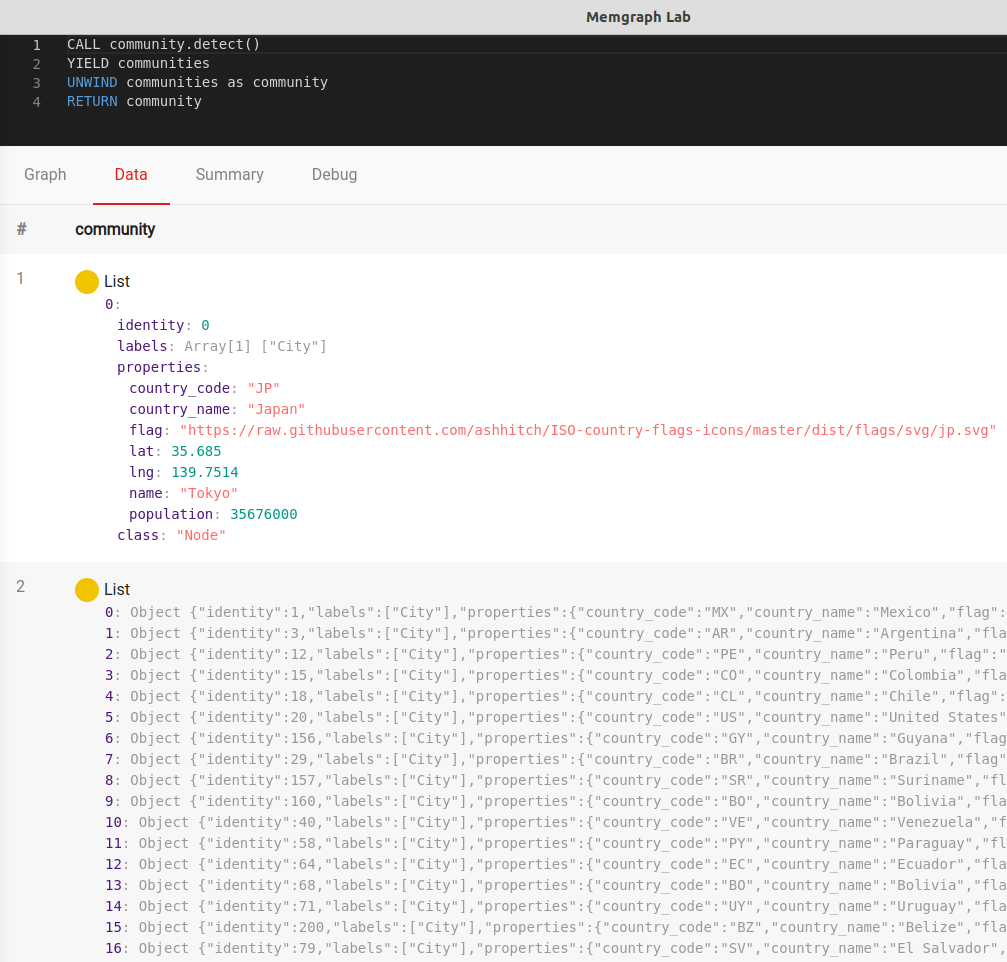
CALL community.detect()
YIELD communities
UNWIND communities as community
RETURN communityThe result is 52 communities with varying numbers of components. Some of the cities in the dataset have been grouped by continent, while some are isolated because of their remote island locations.
Conclusion
Community detection is a powerful tool for graph analysis. From terrorist detection to healthcare initiatives, these algorithms have found their way into many real-world use cases.
The Python NetworkX package offers powerful functionalities when it comes to analyzing graph networks and running complex algorithms like community detection. However, if you're looking to operationalize your graph algorithms and are looking for functionalities such as incremental updates, data persistency, and better performance, you will need to consider using a graph database in conjunction with NetworkX. If you want to find out more about using NetworkX algorithms in Memgraph, read our Reference Guide.